Using Spatial Transcriptomics to Investigate Gene Expression in Cactus Mice
Many of the world’s strangest animals can be found in its harshest environments, where survival in these severe conditions requires exceptional biological adaptations. Nowhere is this more apparent than in deserts, host to extreme daily temperature cycles and a severe lack of water. To overcome the hurdle of water scarcity, some desert animals have evolved the ability to conserve this critical resource. The camel, for example, can quickly rebound from dehydration periods lasting over a week (von Engelhardt et al. 2006). Other organisms find water in more resourceful ways, such as the jackrabbit, which, when standing water is in short supply, stays hydrated by eating moisture-rich plants (Grodsky et al. 2020). But what about other animals missing such built-in defenses—surely all animals still need some form of water intake, right? A mammal able to live its whole life without drinking any water, and without suffering severe organ damage or decrease in functionality, sounds straight out of Star Wars. Yet this creature exists, and it’s no alien!
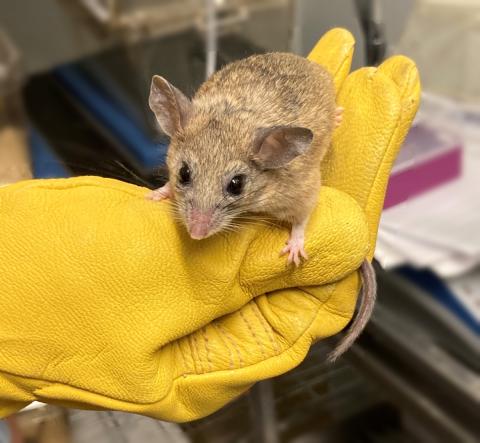

Figure 1: Peromyscus eremicus.
Meet the cactus mouse—Peromyscus eremicus—which, though it will drink water if presented with some, can indeed go its entire life without ever drinking. This species, native to deserts in southwestern North America (Macmillen, 1965), is the focus of Matthew MacManes’ genomics and physiology lab at the University of New Hampshire (UNH). Dr. MacManes, my mentor, is an associate professor in genome enabled biology.
Understanding how these mice accomplish such an incredible feat of retaining full functionality without watermay help scientists develop ways to decrease the negative effects of dehydration in other organisms, including humans. As temperatures continue to rise with global climate change, water will almost certainly become more scarce. Consequently, understanding the methods by which species like the cactus mouse survive without water may help save others from going extinct, or simply help predict which species are likely to withstand climate change the best and which may be in the most danger.
When I heard about Dr. MacManes’ research with cactus mice, I was eager to get involved. I have always loved solving mysteries, from digging into the Washington Post crossword to unraveling plots by Agatha Christie. What puzzle could be more interesting to solve than this seeming exception to the basic rule of life that all animals need water?

Sarah Nicholls
In 2022, I was delighted to receive a Research Experience and Apprenticeship Program (REAP) grant from the Hamel Center for Undergraduate Research, which allowed me to join the lab’s team of graduate students on a project aiming to understand the spatial patterning of gene expression in the kidney and liver—two organs intimately involved in water retention. We planned to accomplish this by creating spatial sequencing libraries, or visual representations of the genes in use, of both organs. We would then overlay these libraries onto high resolution microscope images of the organs themselves to determine the precise location in which each gene was being expressed (in other words, where genes were activated). Throughout the summer and following academic year, I also had the opportunity to collaborate with another graduate student on a project comparing the impacts of high-fat and low-fat diets, as well as free and restricted access to water, on the cactus mouse’s water loss and gene expression.
Background
An organism’s genome, or its full set of genetic material, stores important biological information that serves as the instruction manual for the organism’s survival and growth. The genome is made of DNA, which is itself made of two strands of sequences of biochemical building blocks known as nucleotides. Each individual’s genome serves two important functions: to replicate itself to pass this genetic material on to offspring, and to create proteins, which control many of the biological processes occurring in the organism. In plants, animals, and fungi, DNA is contained within a membrane-enclosed area of the cell, called the nucleus, and is unable to move freely to other parts of the cell.
To begin protein creation, a stretch of DNA known as a gene is transcribed into messenger RNA (mRNA) in a given cell. mRNA is a molecule made of nucleotides like DNA, but it’s much smaller and single-stranded, allowing it to exit the nucleus and have its nucleotide sequence used as a template to build proteins throughout the cell. If many copies of a protein are needed in order to fuel certain biological processes in a given cell, many copies of mRNA molecules are made from the original gene.
In animals, the full genome contains tens of thousands of genes (Cooper, 2000). However, not every protein is needed for each organ’s individual function (think about the differences between your lungs and your brain), so only specific genes are transcribed into RNA in a given cell. Even when the same gene is needed, it may be transcribed at a different frequency than in another cell. This is known as gene expression.
The entirety of an individual’s genome is present in every cell of every tissue regardless of which genes are actually expressed in that cell or tissue, rendering direct DNA analysis unhelpful to examine gene expression. By contrast, the mRNA present in a cell or tissue directly correlates to the genes expressed there, but genomic nucleotide sequences are traditionally recorded in DNA. Therefore, in order to study gene expression, we reverse-engineer from mRNA to DNA in a process known as transcriptomics: examining the mRNA in a given tissue, converting it back to DNA to be sequenced, and thereby determining which genes are operational in that tissue.
While non-spatial transcriptomics tells us which genes are expressed in a given tissue, spatial DNA sequencing, or spatial transcriptomics, is used to determine the precise locations of each gene expressed in that tissue, as tissues are often comprised of different types of cells, each performing their own unique function. If different genes are expressed in different locations, this could indicate that the tissue contains anatomical or functional variation within it (as opposed to homogenous tissue, which is virtually the same throughout), helping contribute to the scientific understanding of the tissue.
Spatial Transcriptomics
For the first phase of my research, we used spatial transcriptomics, which involved the combination of three different fields: a physiological exploration of the cactus mouse; an investigation of its genome via genome sequencing (including spatial gene patterning and libraries); and an analysis of these findings using computation methods. To accomplish this, I learned a variety of technical skills including RNA extraction, techniques in molecular biology, tissue slicing and staining, and microscopy, as well as the biological concepts involved in the process.
Preparing the Kidney and Liver Tissue Slides
To begin, the lab’s transcriptomics team (consisting of myself, Dr. MacManes, and two graduate students, Molly Kephart and Sean Pierre-Louis) met with representatives from the 10X Genomics company, which developed the Visium spatial sequencing technology we used, to better understand the overall spatial transcriptomics process. We then put these concepts into practice. Studying cactus mice from our lab, we collected kidney and liver samples and embedded them in an optimal cutting temperature compound (OCT) using a cold liquid nitrogen-isopentane bath. We then used a cryostat machine at -20℃ to obtain thin, 10-micron sections of the embedded tissue (for reference, 10 microns is much thinner than a strand of hair).
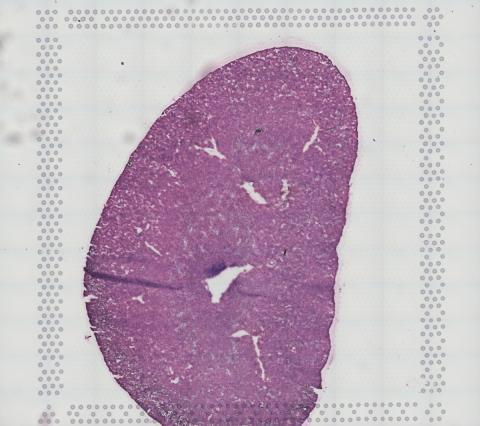
Figure 2: Genome expression slide showing stained kidney sample (photo credit: Molly Kephart).
Two slides were collected from each run: a genome expression slide, containing four slices of organ each from a different mouse, and a tissue optimization slide, containing six slices of organ from a single mouse. The tissue slices were so thin they were barely visible, so we stained them pink on the slides so we could image them clearly on an EVOS microscope managed by UNH’s Center of Integrated Biomedical and Bioengineering Research. We completed multiple practice runs of the above steps before continuing with the experiment to ensure the best possible results.
We used the tissue optimization slide, with the six slices of organ, to determine the optimal length of time of permeabilization. The permeabilization process occurs when an enzyme breaks down cellular membranes, resulting in the organ releasing mRNA like the contents of a popped balloon. We needed to determine the optimal permeabilization time because the genomic libraries we wanted to create are best sequenced with a specific amount of mRNA, and the time of enzyme exposure that releases the optimal amount of mRNA varies among different organs. This meant we needed to individually determine the optimal permeabilization timeframe for both the kidney and the liver.
We first added permeabilization enzyme to each of the six slices of kidney tissue on the same tissue optimization slide for six different lengths of time (3, 6, 12, 18, 24, and 30 minutes). We were able to accomplish these six different times on a
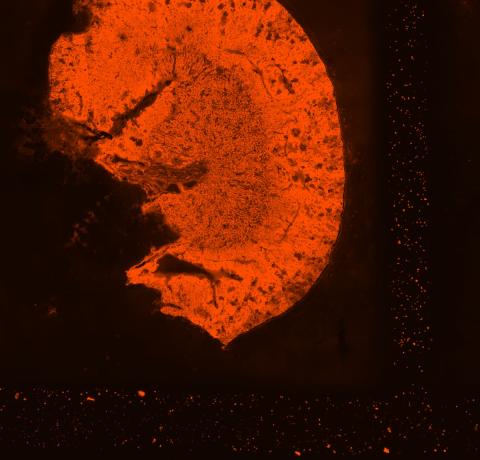
Figure 3: Tissue optimization slide showing fluorescing kidney sample permeabilized for 18 minutes (photo credit: Molly Kephart).
single slide by removing the enzyme from the first sample after three minutes, removing it from the second sample after six minutes, and so on. The tissue optimization slide was useful for this process as it was designed to glow under fluorescent light at the presence of released mRNA. A positive control constructed on the slide contained the enzyme plus previously extracted mRNA, while a negative control contained kidney tissue but no enzyme and therefore didn’t glow at all. When examined under a fluorescent microscope, the slice that glowed the brightest but retained its structural integrity had been permeabilized for the optimal length of time, meaning the optimal amount of mRNA had been released without completely destroying the tissue. We repeated this process for the liver to determine that the optimal permeabilization timeframe for both organs was 18 minutes.
The next step in the experiment was to permeabilize the four slices of tissue on the genome expression slide for the same amount of time in order to obtain the optimal amount of mRNA for sequencing; these four samples were what we used to move forward in the experiment.
Using cDNA to Create Spatial Sequencing Libraries
The genome expression slide contained barcodes, which are very short sequences of nucleotides just like mRNA and DNA, preset in specific locations on the slide. The mRNA released via permeabilization, and therefore genes expressed in the tissue samples, attached to the nucleotides in these barcodes that are engineered to be present in a specific location on the slide. Because the barcodes were made up of a predetermined, known sequence of nucleotides, this would in turn let us locate them during data analysis and determine the precise nucleotide sequences of the mRNA attached to each.
We used the permeabilized mRNA on the genome expression slide and a reverse transcriptase enzyme to create complementary DNA (cDNA) of each mRNA sequence. cDNA simply refers to DNA that has been reverse transcribed from mRNA. Because sequencing data is typically conveyed in terms of DNA, converting back to DNA from mRNA would later allow us to determine the sequence of nucleotides in each original mRNA fragment. After separating the cDNA from the slide, we replicated it using polymerase chain reaction technology (PCR) to increase the quantities of cDNA without changing the ratio of each individual cDNA molecule to other molecules in the sample, then selected a given size of cDNA fragment to sequence and introduced indices. Like the location barcodes, these indices were predetermined nucleotide sequences that would attach to the ends of specific fragments of cDNA, ultimately working with the barcodes to correlate back to the original location of the mRNA on the slide.
Though our work on this project is not yet complete, we have sent the replicated cDNA, with indices attached, to the Hubbard Genome Center next door to our lab to have it sequenced. Our first set of libraries did not result in valid data, and then the cryostat machine broke, but we soldiered on and created a second set. Once the Hubbard Center sends these libraries back this spring, we’ll use bioinformatics to locate our known index and barcode sequences within the libraries, helping us identify the individual cDNA sequences attached to them. Because each barcode corresponds to a specific location within the organ and will therefore determine which mRNA sequences, later converted to cDNA, were found at which location, we’ll be able to overlay the library onto a microscope image of the original organ slice and create a spatial map of gene expression. We’ll also use bioinformatics to determine the levels of gene expression, or the amount of mRNA created, at each specific location within each tissue. With the information from our spatial transcriptomics project, we can begin to understand some of the genetic and molecular processes occurring within the kidney and liver of the cactus mouse.
Non-Spatial Transcriptomics
I was also able to assist with a second project involving transcriptome sequencing of the cactus mouse. This process is similar to the spatial sequencing described above; however, locational barcodes aren’t utilized, so we’re able to correlate the genes expressed only with specific organs instead of specific locations within each organ. Additionally, instead of organ slices, the whole organ is used. I aided Dani Blumstein, a Ph.D. candidate working in the lab, in RNA extractions from the kidneys, liver, hypothalamus, gastrointestinal tract, and lungs of two groups of mice: one that was provided unlimited water, and another that didn’t have access to any water. These five organs were selected for sequencing because each plays a critical role in water retention. As we weren’t investigating the mRNA at specific locations within each organ, we could be less precise with our extractions, using a combination of physical stressors and chemicals (such as TRIzol® and chloroform) instead of permeabilization enzymes to break down the cell membranes and release the mRNA. We prepared replicated cDNA and indices utilizing similar methods as used for our spatial project, and sequenced genomic libraries from this cDNA have recently been returned to the lab by the Hubbard Center. These sequences will allow us to see differences in the overall gene expressions between the mice with water and the mice without water.
This experiment was repeated on a second subset of mice, one on a high-fat diet similar in nutrients to the mice’s lab diet and one on a low-fat diet, to determine the impacts of diet on water retention and gene expression. A prior physiological experiment conducted by the same Ph.D. student suggests that higher-fat diets are correlated with less water loss. The preliminary results from our transcriptomics experiment show that over 4,000 genes in the gastrointestinal tract have different expression across the two treatments, which suggests that diet composition affects the biological processes occurring there. In contrast, fewer than 100 genes in each of the other four organs examined showed different expressions across dietary conditions. This discrepancy in the number of genes being expressed differently suggests that out of these organs, diet has the greatest effect on the gastrointestinal tract. One remaining yet unexplored part of this finding is to understand the role that gut bacteria is playing in generating the differences in gene expression.
More analysis is needed to determine the exact biological processes affected by these differences, which may help us further explain the physiological effects of diet on water retention. Though our study was conducted on the cactus mouse, it’s possible that this is the case in humans as well. For example, Dr. MacManes has been in touch with military scientists to determine if putting troops in drier locations on higher-fat diets would be beneficial. Completing the genomic sequencing to determine how this water retention occurs on a molecular scale will give us a deeper understanding of this process and how it may or may not be beneficial to other species.
What’s Next?
Once our spatial sequencing project is complete, it will inform the upcoming research of the entire MacManes lab. In the immediate future, we plan to compare non-spatial RNA sequences, such as those Dani and I created last summer, to our new spatial maps, allowing us to gain a deeper understanding of what exactly was occurring on the molecular level in previous experiments. The kidney and liver are two organs closely involved with the process of water retention, and no spatial RNA sequencing of any organ in the cactus mouse exists, making our lab the pioneers in this research.
As a long-term goal, we hope to compare these spatial sequences of the kidney and liver of the cactus mouse to sequences from organs of mice that are not desert-adapted, whether in the genus Peromyscus or the genus Mus, the house mouse. Investigating these differences may shed light on what molecular functions permit the cactus mouse to survive at full functionality without water, opening the doorway to future research in low-water tolerance across multiple species. I’m thrilled to continue working with the lab during the next few years as we journey further into this field.
I have been so grateful for the opportunity to research in the MacManes Lab and wanted to extend my gratitude to my collaborators in the lab—Dr. MacManes, Molly Kephart, Sean Pierre-Louis, and Dani Blumstein—for their support, encouragement, and guidance. Thanks as well to the Hamel Center for Undergraduate Research for helping me start my research journey with a REAP grant this past summer. My involvement with this project would not have been possible without the generous financial support of Mr. Dana Hamel, the J.Raymond Hepler Endowed Fund, and Mr.Troy LaPolice; the project would also not have been possible without funding from the National Institutes of Health (NIH). Finally, thanks to my family and friends, who have put up with all my ramblings about RNA and kidneys over the past few months with enthusiasm and patience.
References
Cooper, G.M. 2000. The Cell: A Molecular Approach. 2nd ed. Sunderland (MA): Sinauer Associates.
Grodsky, S.M., L.S. Saul-Gershenez, K.A. Moore-O’Leary, J.P. Whitney, and R.R. Hernandez. 2020. Hare don't care! Consumption of a rare, desert milkweed containing phytochemicals by the black-tailed jackrabbit. Journal of Arid Environments. 174.
Macmillen, R.E. 1965. Aestivation in the cactus mouse, Peromyscus eremicus. Comparative Biochemistry and Physiology. 16(2): 227-248.
von Engelhardt, W., P. Haarmeyer, and M. Lechner-Doll. 2006. Feed intake, forestomach fluid volume, dilution rate and mean retention of fluid in the forestomach during water deprivation and rehydration in camels (Camelus sp.). Comparative Biochemistry and Physiology Part A: Molecular & Integrative Physiology. 143(4): 504-507. https://www.sciencedirect.com/science/article/pii/S1095643306000328.
Author and Mentor Bios
Sarah Nicholls is a biochemistry, molecular and cellular biology major originally from Exeter, New Hampshire. She is in the University Honors Program as well as the Phi Sigma Biological Honor Society, graduating in spring 2025. Sarah’s genomics research on cactus mice in Dr. MacManes’ lab was funded by the Research Experience and Apprenticeship Program (REAP) grant through the Hamel Center for Undergraduate Research. Her lifelong fascination with solving puzzles drew her to genomics research, which she sees as a process of decoding the secrets of life from a tiny molecule. The species of the cactus mouse, a mammal that does not need water to survive, seemed like the ultimate genomic puzzle to solve. Through this research, Sarah also gained a deeper knowledge of the research process, with all the setbacks and problem-solving that occurs throughout. While graduation is a couple years away, she plans to continue research in pursuing a doctorate in either genetics or biochemistry.
Dr. Matthew MacManes is an associate professor of genome enabled biology in the Molecular, Cellular, and Biomedical Sciences, joining UNH in 2013. The MacManes lab is interested in understanding how adaptive evolution has shaped animal diversity through studying physiology and genomics. One of several ongoing projects is understanding how desert mammals survive without water. This is the research that Sarah Nicholls participated in, beginning in the summer of 2022 through the Research Experience and Apprenticeship Program (REAP), which then continued into the academic year.
Copyright 2023, Sarah Nicholls